Gervaise Henry
Scan contact card
Building data & AI platforms for genomics and multi-omics research. VP, Forward Deployed Engineering at Manifold — bench and computational biologist turned platform leader.
I’ve spent my career at the intersection of biology, data, and infrastructure: starting at the bench running high-throughput screens and single-cell RNA-seq, then moving into computational biology, cloud architecture, and ultimately leading the engineering organizations that build platforms for genomics and multi-omics research at scale. I’ve built and managed global teams of solutions architects, bioinformatics engineers, and services engineers across the US, Europe, and Asia. AWS Certified Solutions Architect & Cloud Practitioner. Certified Scrum Product Owner.
Skills
- Solutions Engineering Leadership
- Genomics & Multi-Omics Platforms
- Cloud Architecture (AWS)
- AWS Certified Solutions Architect - Associate
- AWS Certified Cloud Practitioner
- Scrum Alliance Certified Scrum Product Owner
- R
- Python
- Nextflow
- WDL
- git
- linux
- bash
- Static Site Generators
- Serverless Deployment
- GitLab Pages
- html5
- css3
- javascript
- amazonwebservices
- Azure
- googlecloud
- S3
- EC2
- Batch
- Lambda
- DynamoDB
- API Gateway
- Certificate Manager
- Route53
- CloudFront
- CloudTrail
- Athena
- Docker
- Singularity
- Bulk RNA-seq
- Gene Expression Microarray
- scRNA-seq
- scTCR-seq
- Spatial Transcriptomics
- Variant Calling
- Proteomics (Mass Spec)
- Molecular Biology Techniques
Experience
VP, Forward Deployed Engineering
Lead Forward Deployed Engineering at Manifold, the enterprise agent platform for life sciences — responsible for deploying specialist agents into customer environments and owning post-sales technical delivery while staying active in pre-sales.
- Building and scaling the Forward Deployed Engineering organization, with direct reports spanning FDEs and Forward Deployed Builders, and expanding the team to meet growing customer demand for agent deployment and implementation.
- Provide technical leadership for Forward Deployed Engineers across the broader organization, guiding how they engage with customers on complex implementations even where reporting lines sit elsewhere.
- Partner with the pre-sales Solutions team across the technical customer journey — contributing to discovery, scoping, proof-of-concept development, and platform demonstrations — while owning post-sales implementation, technical delivery, and ongoing customer partnership.
- Maintain broad coverage across Manifold’s customer base with concentrated focus on strategic accounts — particularly high-risk, high-reward engagements where customer problems are technically complex and platform fit requires deep scientific and architectural collaboration.
- Build proof-of-concept, reference implementation, and demo assets across three purposes: supporting pre-sales in customer acquisition, driving expansion within existing accounts, and extending platform functionality for broader customer applicability.
- Partner with Product and Engineering to translate field signal into roadmap priorities and to stress-test new capabilities against customer needs before they ship.
- Serve as a liaison between customers, sales, product, engineering, and support, ensuring customer technical and scientific needs are represented throughout the organization.
VP, Head of Data Engineering
Led data infrastructure and management strategy for Champions’ research and development efforts, with responsibility for the data platform supporting multi-omics and research data workflows in alignment with FAIR principles.
- Set direction on modernizing Champions’ analytic pipelines, initiating rebuilds to bring them in line with current standards for reproducibility, maintainability, and scalability.
- Migrated the analytic compute environment onto a foundation that delivered meaningful reductions in pipeline runtime and cost, improving the economics of recurring analyses.
- Initiated the build-out of data provenance, lineage, and observability infrastructure — evaluating tooling and beginning implementation of the governance layer needed to support regulated multi-omics and research data at scale.
- Owned data organization and deployment for new data commercialization efforts, structuring datasets for external availability and partnering with business stakeholders to bring data products to market.
- Supported sales efforts as the senior scientific and technical voice, providing subject-matter expertise in customer conversations and shaping the technical positioning of data offerings.
- Contributed to new assay development efforts, particularly on the analytical and data-handling side — advising on how downstream analysis requirements should shape assay design, data capture, and output structure.
- Served as the senior technical voice for data strategy, partnering with R&D and executive leadership on platform direction, tooling selection, and long-term data architecture decisions.
Head of Professional Services Engineering
Led the Professional Services Engineering organization — a global team of 15–20 architects, bioinformatics engineers, services engineers, and contractors across the US, Europe, and Asia — responsible for all technical services delivery on the DNAnexus platform.
- Expanded scope from leading the Solutions Engineering function to leading all technical staff across Professional Services, following consistent delivery improvements under prior management.
- Oversaw services delivery across a global, distributed team — including customer-facing engineering, platform integrations, custom solution development, and data engineering for customer data ingestion.
- Managed a sizeable offshore contractor contingent alongside full-time staff, balancing delivery throughput with margin discipline as the services function reached its highest profitability during this period amid continued scaling.
- Partnered with Product Engineering and Product Management on roadmap priorities informed by customer delivery experience, and built proof-of-concept assets to support architectural decisions on new platform capabilities.
- Served as a technical liaison between sales, customers, and internal product and support teams — deployed frequently to strategic or at-risk customer engagements to resolve escalations and restore delivery trajectory.
Manager, Solutions Science Engineering
Managed the Solutions Engineering organization through a period of leadership transition, expanding scope mid-tenure to include a subset of the Professional Services engineering team.
- Led a team of 5–6 Solutions Engineers distributed across the US and EMEA, supporting presales engagements globally across all customer verticals.
- Took on additional management responsibility for part of the Professional Services engineering function following a leadership vacuum, tightening integration between presales and services — notably in SOW scoping accuracy and services utilization — which led to subsequent promotion to Head of PS Engineering.
- Built DNAnexus’s external technical presence in Europe and North America, delivering invited oral presentations at EU Biobank Week 2024, ACMG 2024, BioTechX Europe 2023, and the Precision Medicine Leaders Summit 2023, with additional panel participation at BioIT World 2023 on precision medicine and FAIR multimodal data.
- Built and refined proof-of-concept and demo assets to demonstrate platform capabilities against customer-specific scientific and technical needs.
- Partnered with Product Engineering and Product Management on roadmap prioritization based on field signal, and served as liaison between sales, customers, product, professional services, and support.
Senior Solutions Engineer
Individual contributor presales engineer supporting customers globally across all verticals, with particular depth in top-5 pharma and diagnostics.
- Partnered with sales on customer discovery, scoping, and technical qualification — translating customer scientific and operational needs into platform-fit assessments and architectural proposals.
- Built proof-of-concept assets to validate the DNAnexus platform against real customer workflows, including substantial WDL and Nextflow pipeline development and lift-and-shift work.
- Delivered platform demonstrations, technical training, and white-glove technical support for strategic accounts throughout the presales cycle.
- Served as technical liaison between sales, customers, and internal product, professional services, and support teams — recognized for technical depth and customer trust that led to first-line management responsibility following departures of other team leaders.
Computational Biologist
Applied computational, cellular, and molecular techniques to investigate normal biology of the lower urinary tract and the development and progression of benign and malignant pathologies. Focused on single-cell and bulk RNA-sequencing of FACS-sorted populations to characterize normal and diseased cellular heterogeneity. Developed and deployed standardized pipelines for data analysis and exploration, including cloud-based (AWS, Azure) implementations.
Research Associate
Designed, conducted, and analyzed cellular and molecular experiments investigating normal prostate biology and the development and progression of benign prostatic hyperplasia and prostate cancer. Leveraged single-cell and bulk RNA-sequencing of FACS-sorted populations to characterize cellular heterogeneity in healthy and diseased tissue, and identified immune infiltrate signatures in primary human prostate samples to associate with cell-type-specific gene expression, epithelial dynamics, and clinical outcomes. First or co-authored eight journal publications during this period, including a first-author paper in Cell Reports (2018).
Research Associate
Developed high-content image-based assays for high throughput drug discovery. Optimized high throughput assays for early-stage drug discovery on marine natural products.
RA/TA
Used cellular and molecular biology techniques to investigate the effect of adipocytes and obesity on the multiple myeloma bone marrow tumor microenvironment. Techniques included cell culture (including co-culture), immunoblotting, ELISA, qPCR, tubulogenesis assays, invasion assays, viability assays, immunofluorescence, and confocal microscopy. Independently taught undergraduate lab courses, including design of laboratory content, assessments, grading, and student counseling.
Courses taught: Bio 200 Structure and Function of the Human Body, Bio 210 General Biology II, Bio 300 Cell Biology, Bio 356 Genetics.
Lab Technician
Investigated the relationship between the peptidyl-prolyl cis/trans isomerase Pin1 and IL13 in asthma models, using cellular, molecular, and biochemical techniques. Managed lab budget, ordering, and vendor relationships.
Screening Scientist
Conducted high-throughput compound and RNAi screens in the HTS/RNAi Core, working with a 200,000-compound library, whole-genome human siRNA, Drosophila dsRNA, and custom libraries. Primary project used compound and RNAi screens to functionally categorize NSCLC cell lines into familial groups, identifying molecular susceptibilities exploitable based on biomarkers. Contributed across all aspects of projects: assay choice and development, optimization, implementation, data analysis, bioinformatics, and follow-up experiments. Additional projects included screens for osteoclast differentiation modifiers, novel-mechanism antimicrobial agents, iron homeostasis regulators (siRNA), and miRNA sensitizers to sub-lethal chemotherapy in cancer cells with specific genetic profiles.
Summer Undergraduate Research Intern
Screened for NF-κB inhibitors using a cell-based luciferase reporter assay on a library of partially purified marine natural products. Confirmed prospective inhibitors via flow cytometry and immunoblotting, and identified active compounds using HPLC, MS, and 1D/2D NMR.
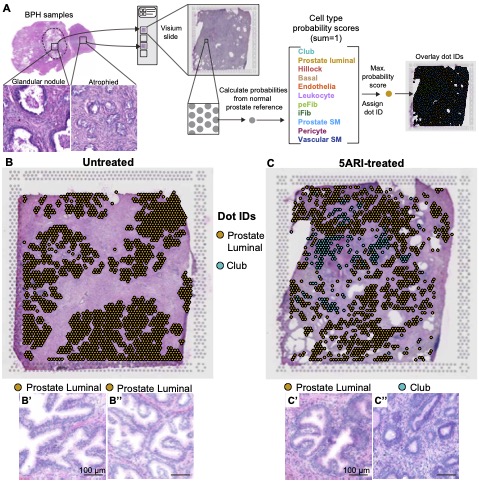
The Journal of Pathology - 5-alpha reductase inhibitors induce a prostate luminal to club cell transition in human benign prostatic hyperplasia
Joseph DB, Henry GH, Malewska A, Reese JC, Mauck RJ, Gahan JC, Hutchinson RC, Mohler JL, Roehrborn CG, Strand DW
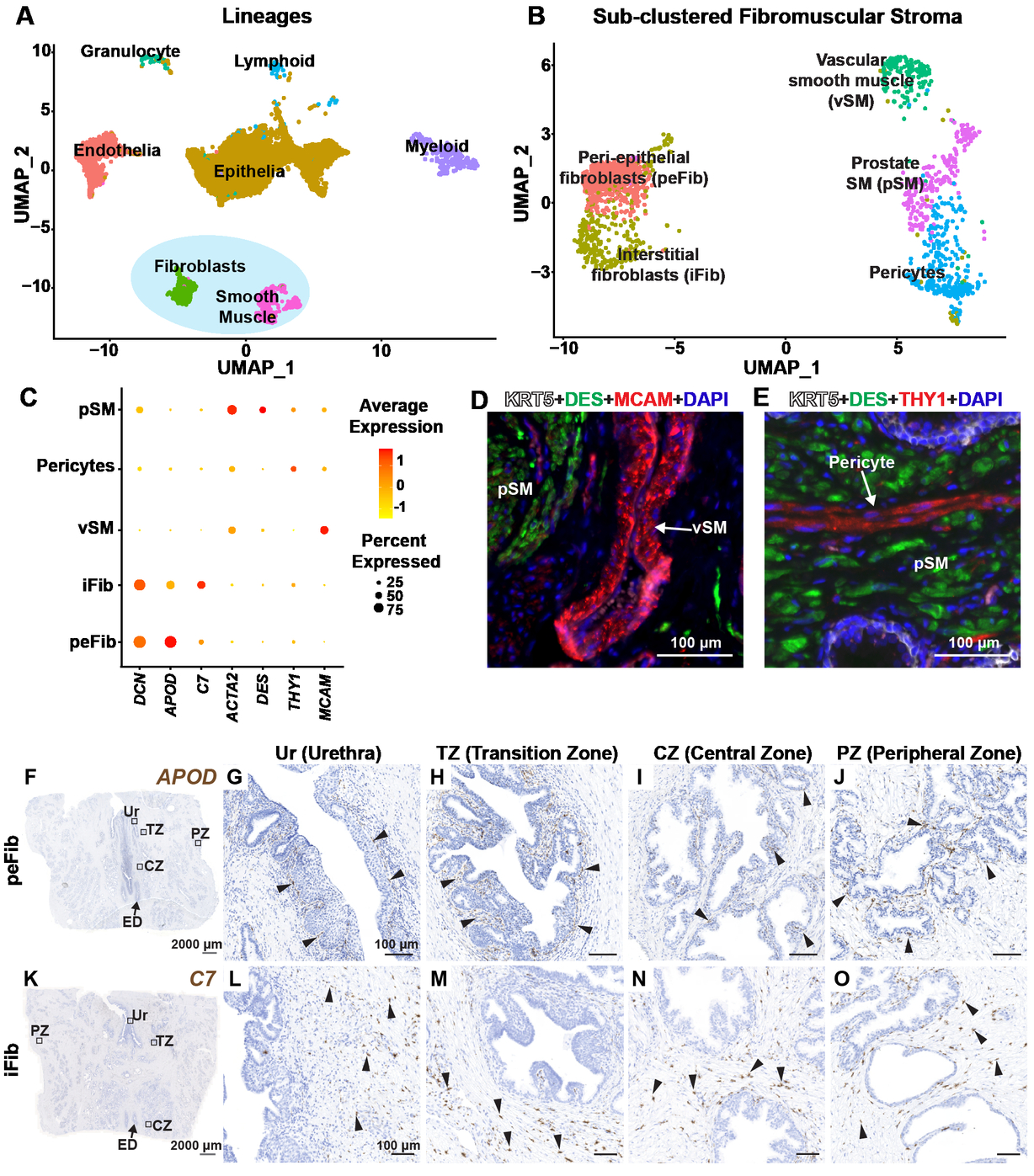
The Journal of Pathology - Single cell analysis of mouse and human prostate reveals novel fibroblasts with specialized distribution and microenvironment interactions
Joseph DB, Henry GH, Malewska A, Reese JC, Mauck RJ, Gahan JC, Hutchinson RC, Malladi VS, Roehrborn CG, Vezina CM, Strand DW
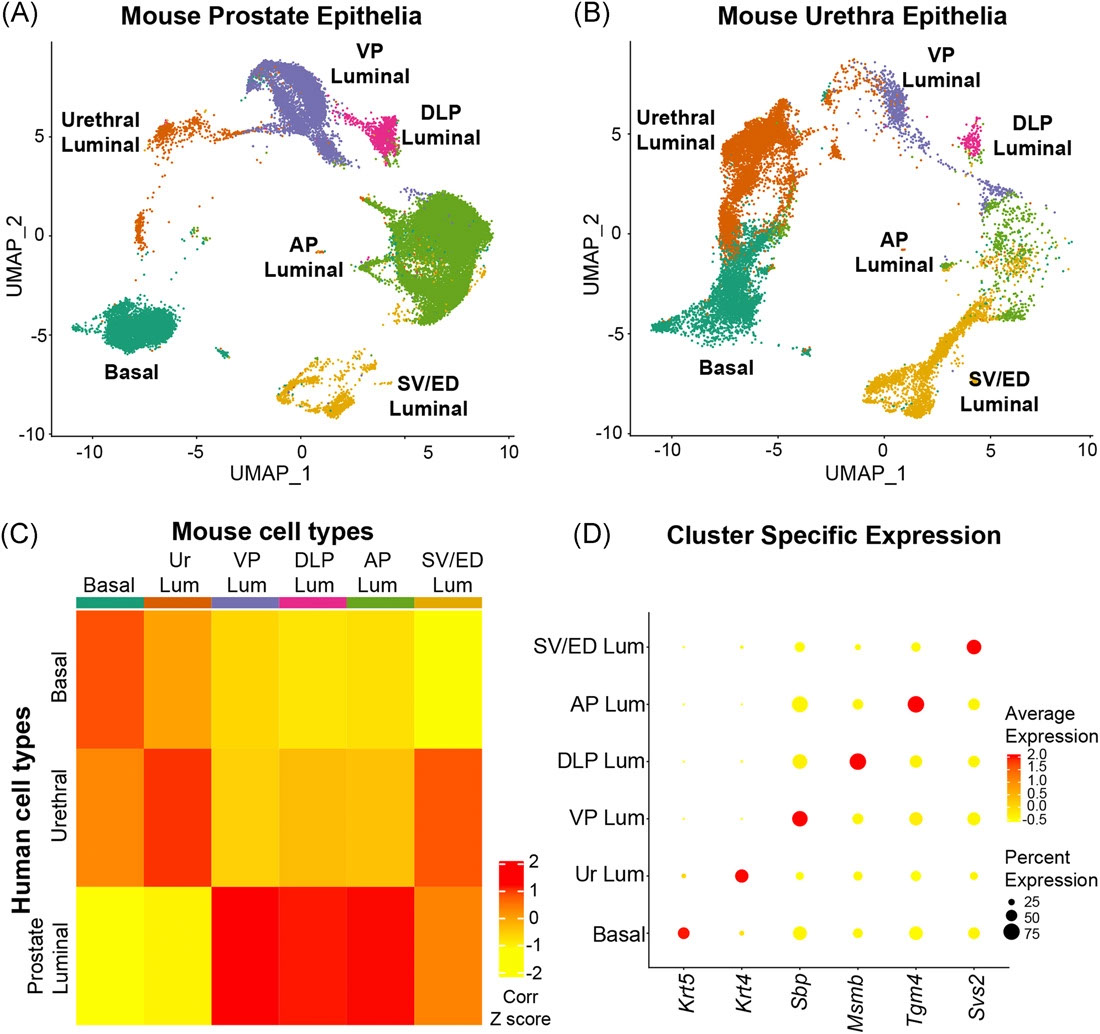
The Prostate - Urethral luminal epithelia are castration-insensitive cells of the proximal prostate
Joseph DB, Henry GH, Malewska A, Iqbal NS, Ruetten HM, Turco AE, Abler LL, Sandhu SK, Cadena MK, Malladi VS, Reese JC, Mauck RJ, Gahan JC, Hutchinson RC, Roehrborn CG, Baker LA, Vezina CM, Strand DW
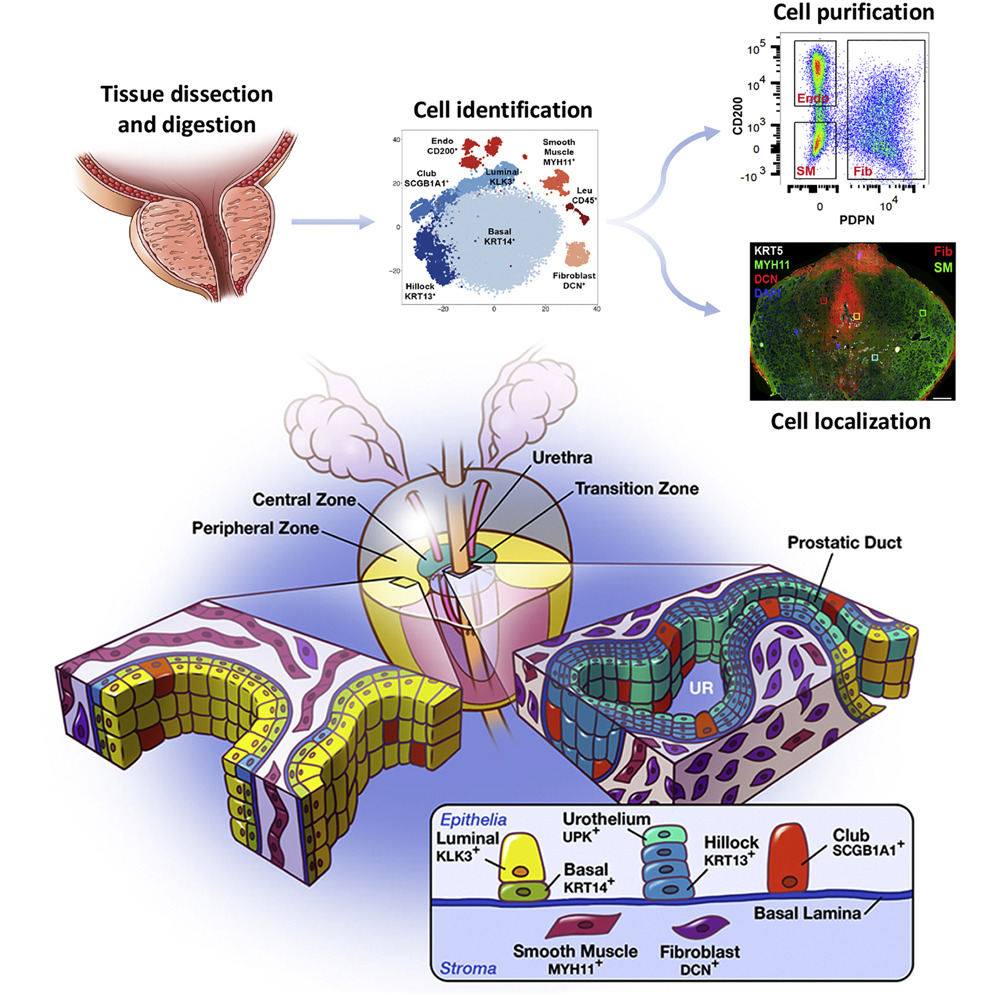
Cell Reports - A cellular anatomy of the normal human prostate
Henry GH, Malewska A, Joseph DB, Malladi VS, Lee J, Torrealba J, Mauck RJ, Gahan JC, Raj GV, Roehrborn CG, Hon GC, MacConmara MP, Reese JC, Hutchinson RC, Vezina CM, Strand DW
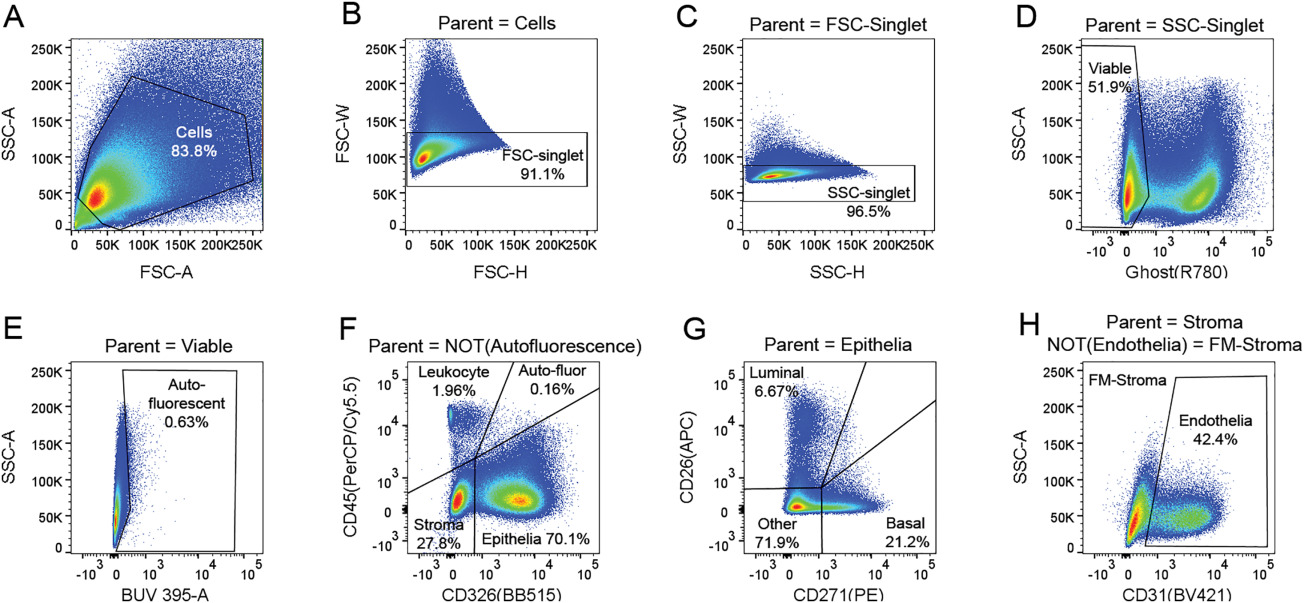
Cytometry Part A - OMIP-040: Optimized gating of human prostate cellular subpopulations
Henry GH, Loof N, Strand DW
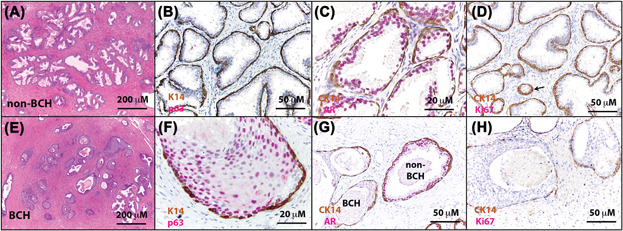
The Prostate - Molecular pathogenesis of human prostate basal cell hyperplasia
Henry GH, Malewska A, Hutchinson R, Gahan J, Mauck R, Francis F, Torrealba J, Roehrborn C, Strand DW
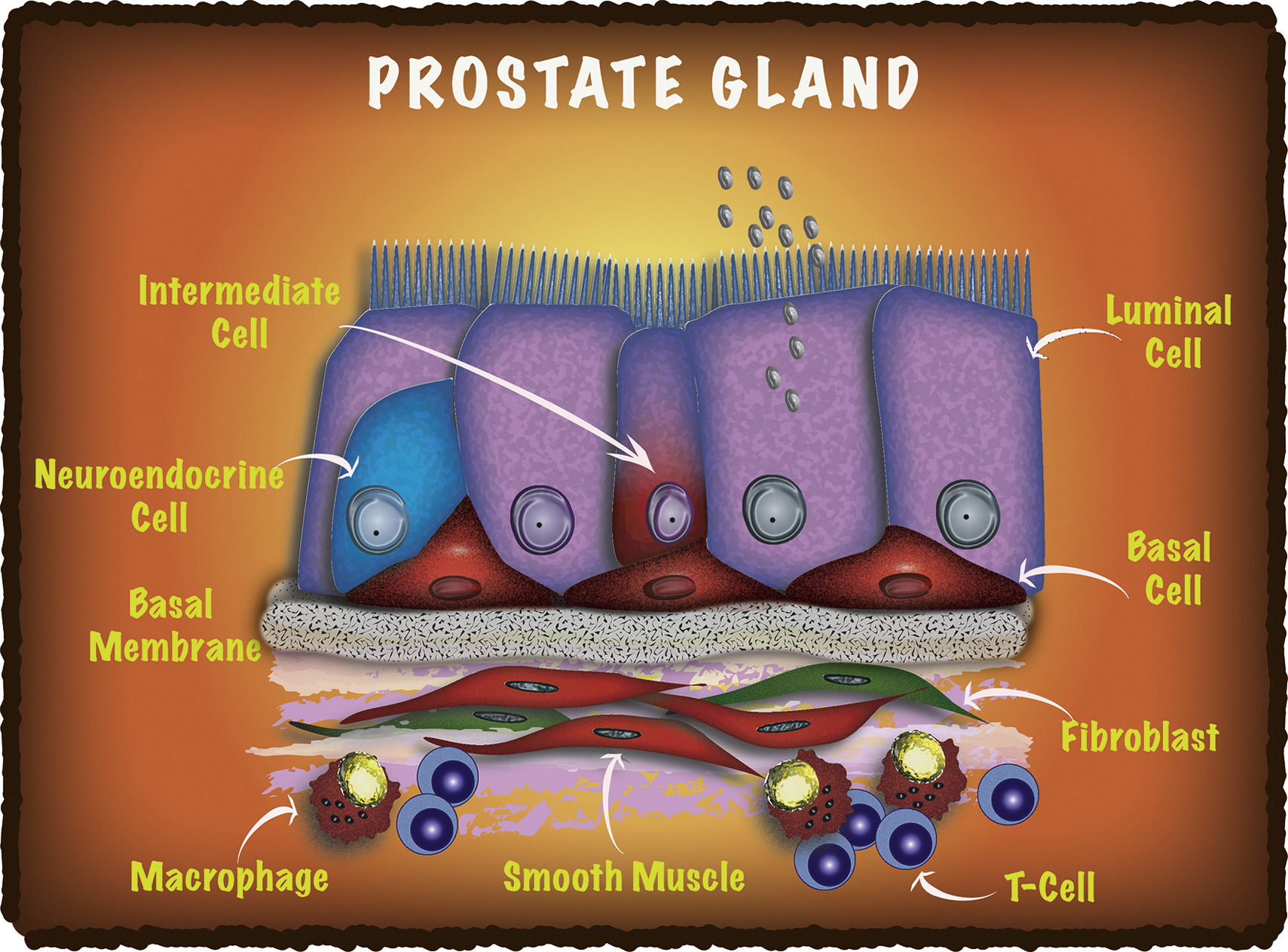
Differentiation - Isolation and analysis of discrete human prostate cellular populations
Strand D W, Aaron L, Henry G, Franco O E, Hayward SW
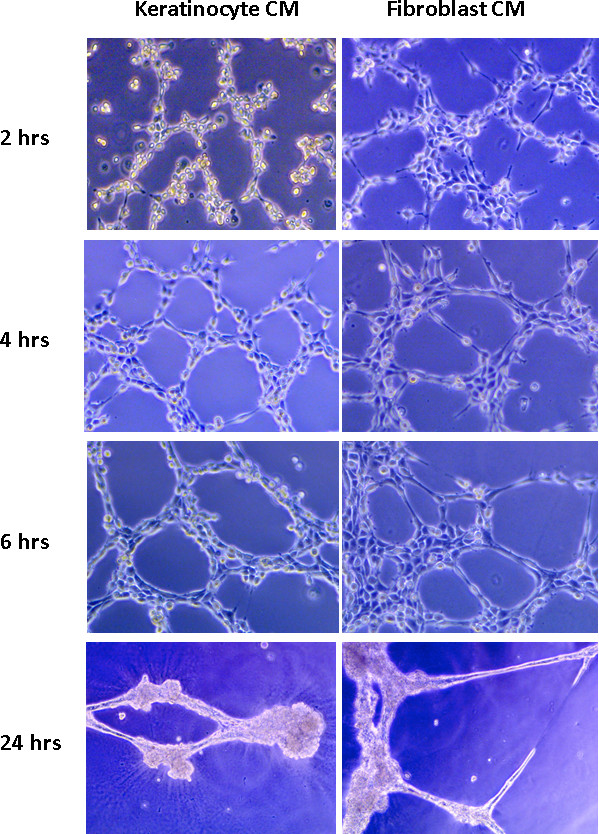
JoVE - Endothelial cell tube formation assay for the in vitro study of angiogenesis
DeCicco-Skinner KL, Henry GH, Cataisson C, Tabib T, Gwilliam JC, Watson NJ, Bullwinkle EM, Falkenburg L, O’Neill RC, Morin A, Wiest JS
NextGen Omics, Spatial & Data US - March 2026
From Spatial Transcriptomics to Multimodal Discovery: Scaling AI Driven Genomics on Manifold
Mohta V, Farhi S, Henry GH
EU Biobank Week - April 2024
Efficiently Analyzing Population-Scale Genomics Data
Sedlakova, A, Henry GH
ACMG - March 2024
Automated Genome Interpretation Solution to Accelerate Diagnostic Decisions via Intelliseq GeneSpect™ Reporter
Szklarczyk-Smolana K, Pacewicz K, Henry GH
BioTechX Europe - October 2023
Bridging the Gap: Solutions for Data Integration, Security, and Collaborative Exploration in Precision Medicine
Rao A, Henry GH
Precision Medicine Leaders Summit: Advances in Single Cell & Spatial Biology - September 2023
Democratizing Access to Single-Cell RNA Sequencing with the Parse Biosciences Evercode Platform
Henry GH, Roco C
Precision Medicine Leaders Summit: Advances in Single Cell & Spatial Biology - September 2023
Innovation Panel: Building the Spatial Biology Ecosystem
Enderlein C, Clark JN, Emanuel G, Henry GH, Kim J
BioIT World - May 2023
Towards Precision Medicine: Datasets, Computation, and Data Integration for Patient Subsetting Research - Part I
Busby BR, Chu E, Floden E, Henry GH, Huitt E
BioIT World - May 2023
Achieving FAIRness with Multimodal Data
Busby BR, Dawson E, Gillis M, Henry GH, Holt R, Neilley V
Cold Spring Harbor: Single Cell Analysis - November 2019
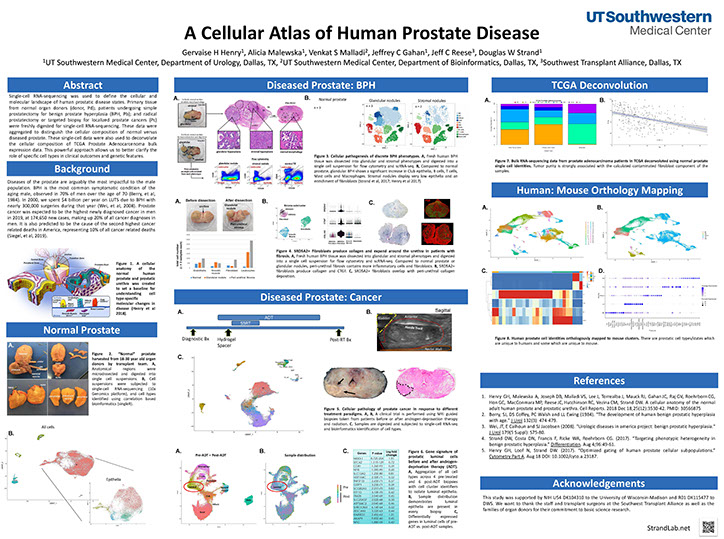
A Cellular Atlas of Human Prostate Disease
Henry GH, Malewska A, Malladi VS, Lee J, Gahan JC, Reese J, Strand DW
American Urological Association: Annual Meeting - May 2019
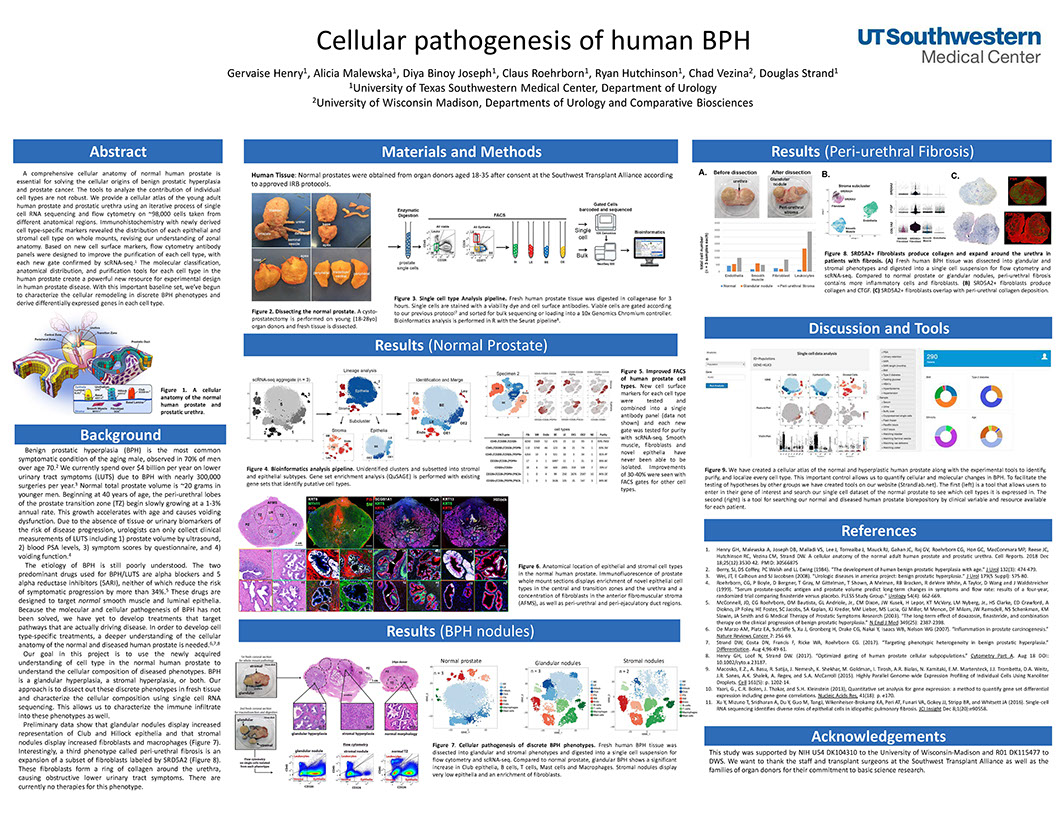
Cellular pathogenesis of human BPH
Henry G, Malewska A, Binoy Joseph D, Roehrborn C, Hutchinson R, Vezina C, Strand D
Collaborating for the Advancement of Interdisciplinary Research in Benign Urology: Meeting - December 2018
Building a comprehensive cellular anatomy of the normal and diseased human prostateia
Henry GH, Malewska A, Joseph DB, Malladi VS, Lee J, Torrealba J, Mauck RJ, Gahan JC, Raj JV, Roehrborn CG, Hon GC, MacConmara MP, Reese JC, Hutchinson RC, Vezina CM, Strand DW
Society for Basic Urologic Research: Annual Meeting - November 2018
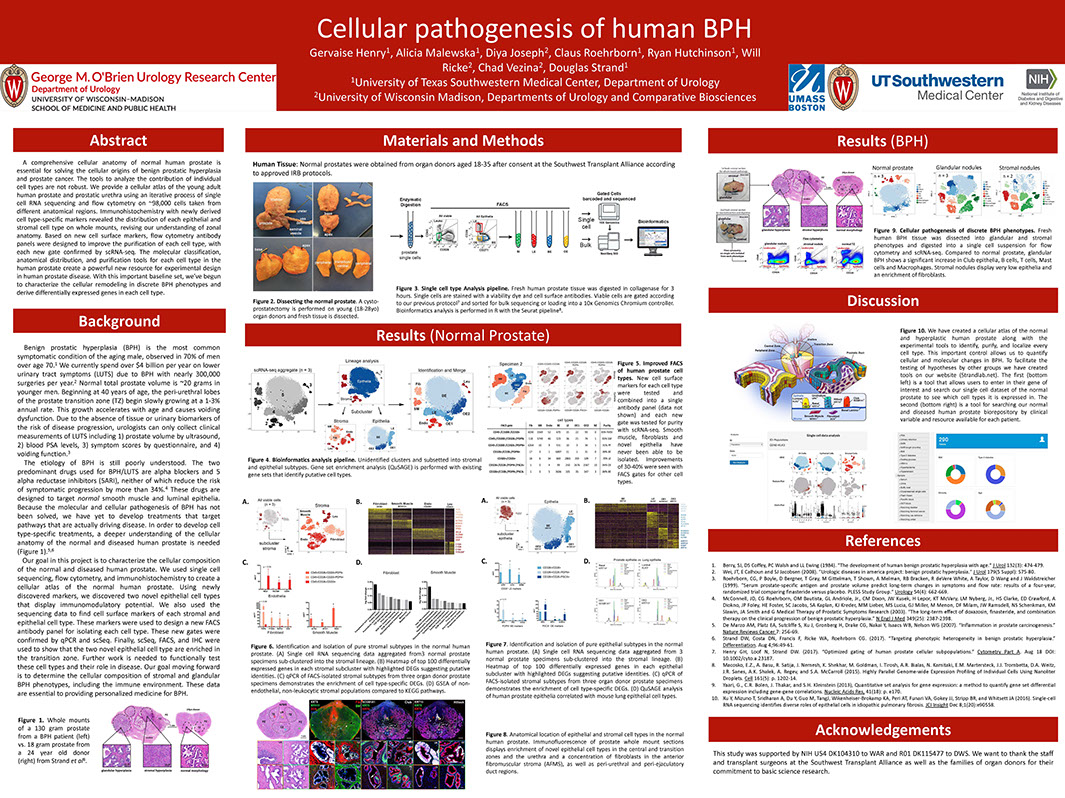
A Cellular Anatomy of the Normal Human Prostate and BPH
Henry G, Malewska A, Binoy Joseph D, Malladi V, Lee J, Gahan J, Mauck R, Raj G, Roehrborn C, Hon G, Reese J, Hutchinson R, Vezina C, Strand D
10x Genomics User Group Meeting: Houston - May 2018
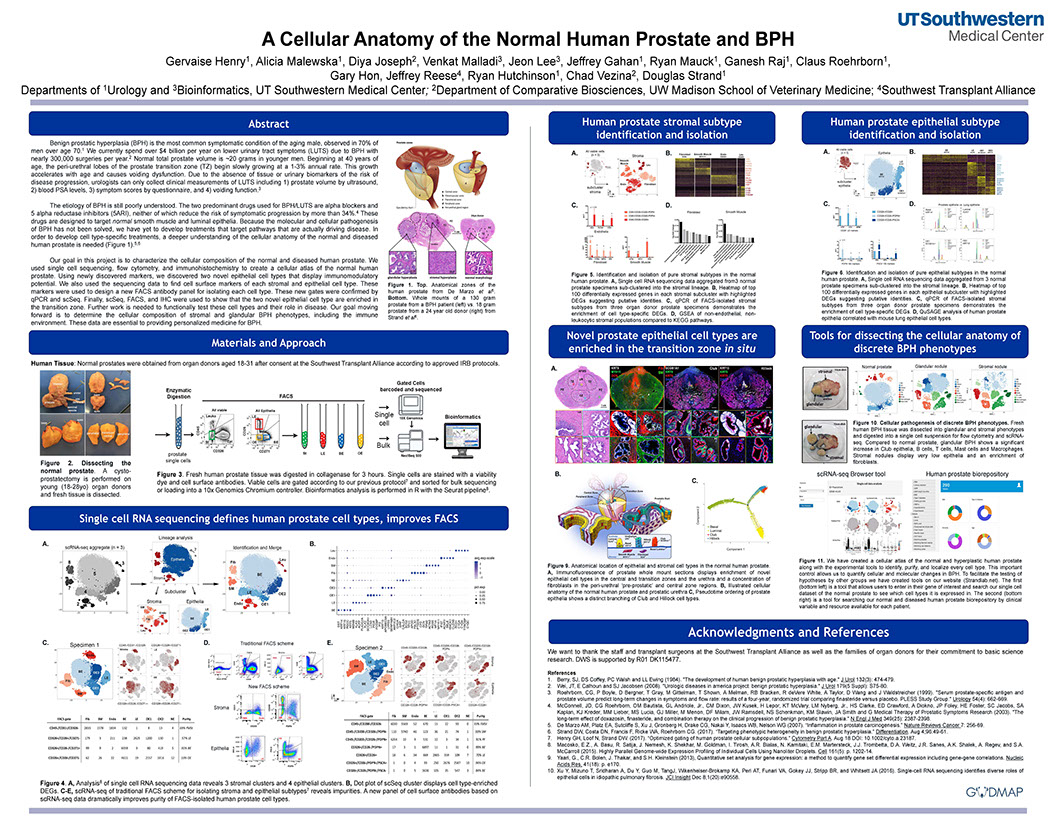
Determining Cellular Heterogeneity in Human Prostate using the Chromium™ Single Cell 3’ Solution
Henry GH
FlowTex Conference - February 2018
Determining cellular heterogeneity in human prostate with flow cytometry and single-cell RNA-sequencing
Henry GH
North Texas Flow Cytometry Conference - December 2017
Determining cellular heterogeneity in human prostate with flow cytometry and single-cell RNA-sequencing
Henry GH
Society for Basic Urologic Research: Annual Meeting - November 2017
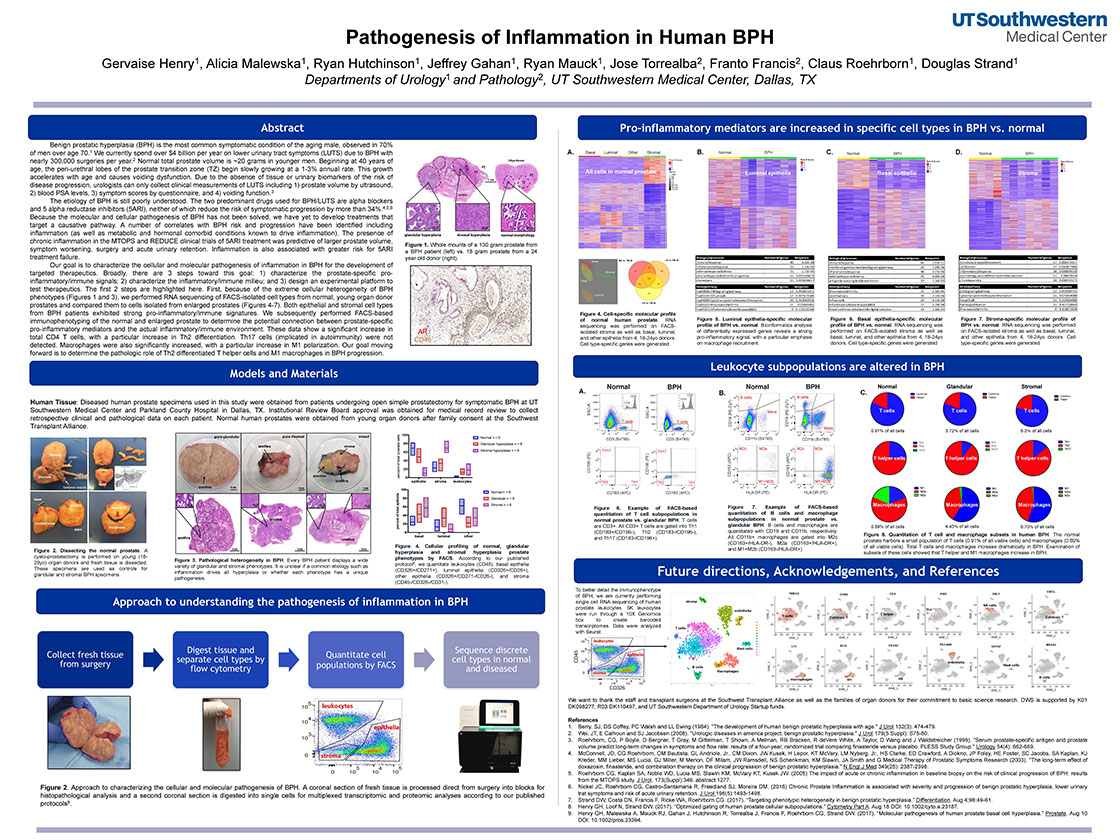
Pathogenesis of Inflammation in Human BPH
Henry G, Malewska A, Hutchinson R, Gahan J, Mauck R, Torrealba J, Francis F, Roehrborn C, Strand D
North Texas Flow Cytometry Conference - December 2016
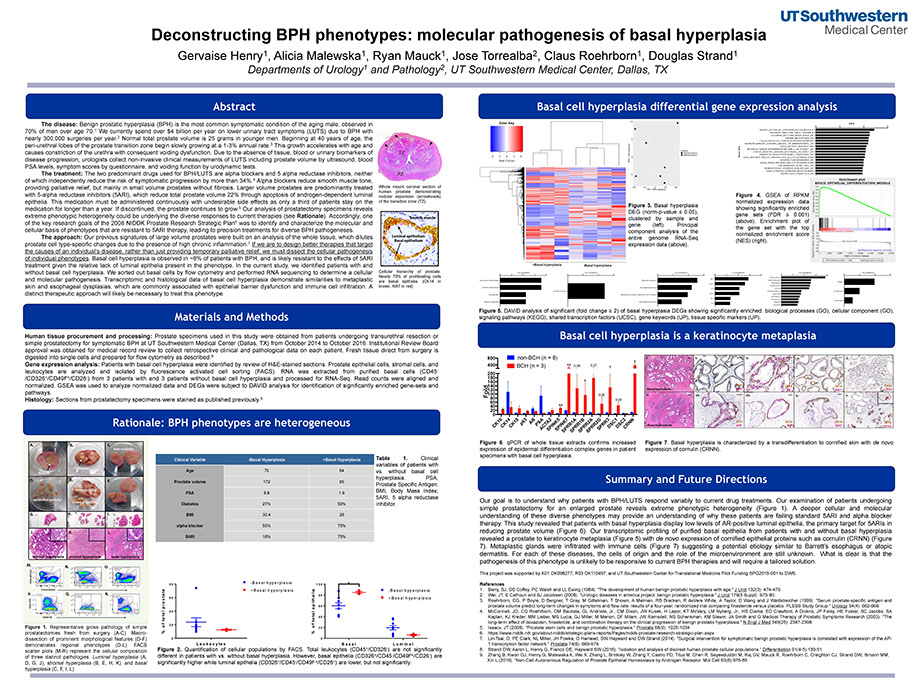
Deconstructing BPH Phenotypes: Molecular Pathogenesis of Basal Hyperplasia
Henry GH
Society for Basic Urologic Research: Annual Meeting - November 2016
Deconstructing BPH phenotypes
Henry G, Malewska A, Mauck R, Torrealba J, Roehrborn C, Strand D

Society for Basic Urologic Research: Annual Meeting - November 2015
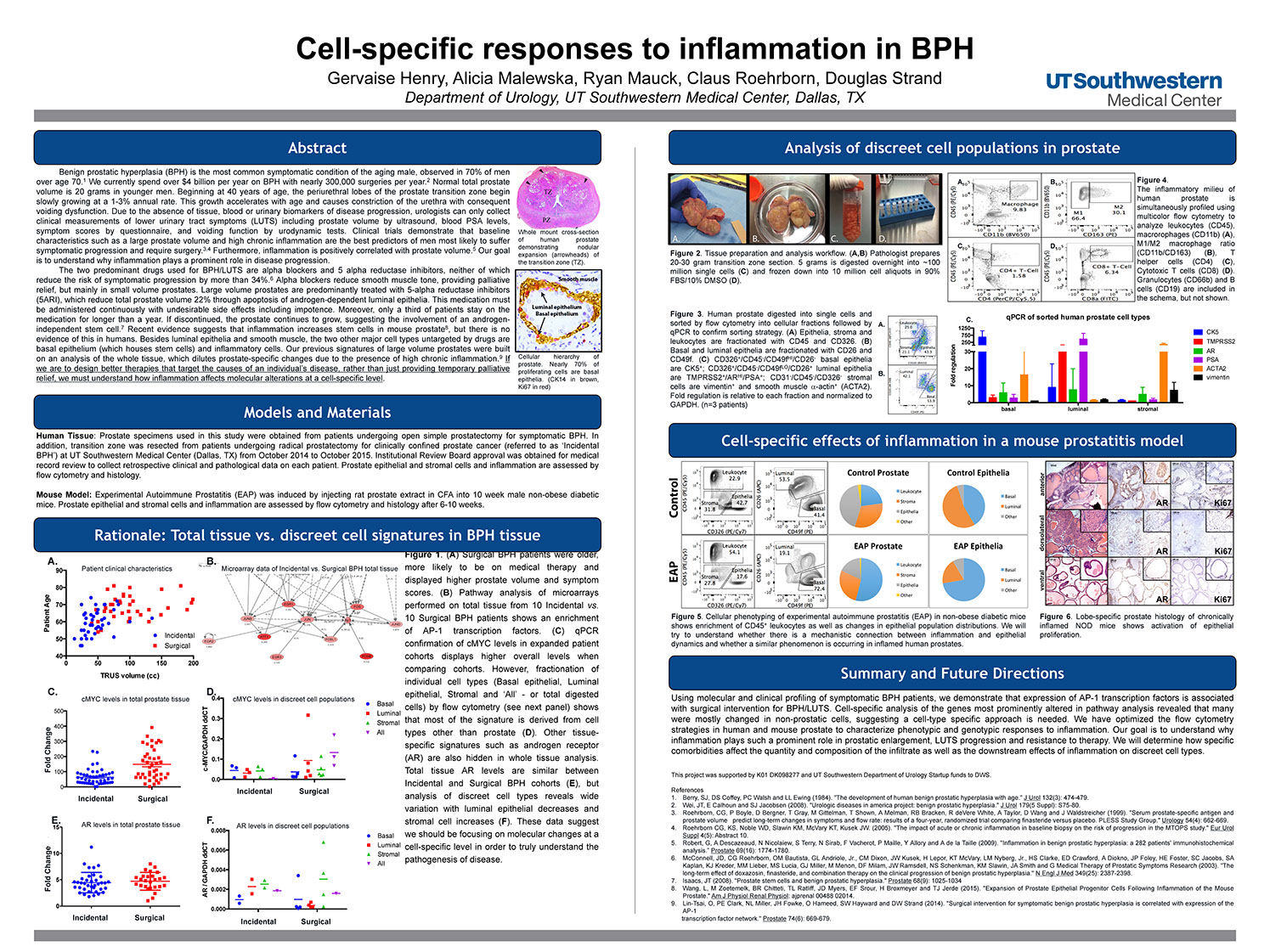
Cell-specific responses to inflammation in BPH
Henry G, Malewska A, Mauck R, Roerborn C, Strand D

American Association of Cancer Research: Annual Meeting - April 2014
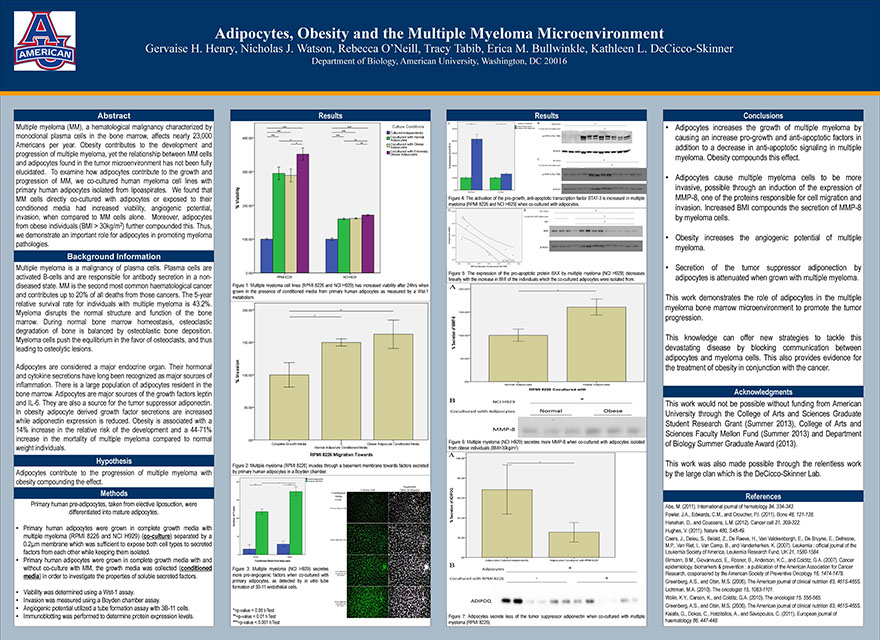
Adipocytes, Obesity and the Multiple Myeloma Microenvironment
Henry GH, Watson NJ, O’Neill R, Tabib T, Bullwinkle EM, DeCicco-Skinner KL
Open Source
A collection of open source visualizations, code, and data that I created or contributed to.
cellxgene: Single cell analysis of mouse and human prostate reveals novel fibroblasts with specialized distribution and microenvironment interactions

Single-cell RNA-sequencing of adult human prostates and urethras from organ donors, and BPH (glandular and stromal) patients as well as adult mouse lower urinary tracts.
RNA-seq Analytic Pipeline for GUDMAP/RBK

This pipeline conducts end-to-end RNA-seq analysis on replicates stored in the GenitoUrinary Development Molecular Anatomy Project and ReBuilding the Kidney consortium. This pipeline was created in a collaboration between the Strand Lab and Bioinformatics Core Facility (BICF) at UT Southwestern Medical Center.This pipeline conducts end-to-end RNA-seq analysis on replicates stored in the GenitoUrinary Development Molecular Anatomy Project and ReBuilding the Kidney consortium. This pipeline was created in a collaboration between the Strand Lab and Bioinformatics Core Facility (BICF) at UT Southwestern Medical Center.
Strand Lab analysis of single-cell RNA sequencing
This code was used for the single-cell RNA-sequencing analysis in the Strand Lab at UT Southwestern Medical Center
BICF Cellranger count Analysis Workflow

BICF Cellranger count Analysis Workflow is a wrapper for the cellranger count tool from 10x Genomics. This pipeline, used by the Bioinformatics Core Facility at UT Southwestern, takes fastq files from 10x Genomics single-cell gene expression libraries and passes them to cellranger count, managing parallelization of multiple runs, as well as, aggregation as appropriate. This pipeline is primarily used with a SLURM cluster on BioHPC, but it should be able to run on any system that Nextflow supports. Additionally, this pipeline is designed to work using a simple web interface.
GEO: Human prostate luminal epithelia adopt a club-like state in response to low androgen levels

Spatial transcriptomics of treatment naive and 5-alpha reductase inhibitior treated human BPH tissue
GEO: Single cell RNA-sequencing of verumontanum region and peripheral zone of normal human prostate

Dissected verumontanum region and peripheral zone region from a normal human prostate sample were digested into single-cell suspensions, cells were subject to single-cell RNA-sequencing using the 10x Genomics platform
GEO: Single-cell RNA-sequencing of adult human normal and BPH (glandular and stroma) prostates and urethra

Single-cell RNA-sequencing of adult human prostates and urethras from organ donors, and BPH (glandular and stromal) patients.
GEO: Single-cell RNA-sequencing of adult mouse lower urinary tracts v2

Single-cell RNA-sequencing of adult mouse prostates and urethras.
GUDMAP: Single cell analysis of mouse and human prostate reveals novel fibroblasts with specialized distribution and microenvironment interactions

Sequencing data relating to the in preparation publication: 'Single cell analysis of mouse and human prostate reveals novel fibroblasts with specialized distribution and microenvironment interactions'
